10 Mar 2025 | Snakemake Hackathon

Jens and Johanna attended the Snakemake Hackathon at the Cern in Geneva.
The Hackathon ended with the release of Snakemake 9.0.

The Algorithmic Bioinformatics group at Saarland University is headed by Prof. Sven Rahmann. It belongs to both the Mathematics and Informatics Faculty (“MI”) and the Center for Bioinformatics (ZBI) and is part of Saarland Informatics Campus (SIC). Our research focuses on method development (algorithms and data structures) for concrete problems that arise in biological data analysis. We mainly teach in the Bioinformatics degree programs.

Jens and Johanna attended the Snakemake Hackathon at the Cern in Geneva.
The Hackathon ended with the release of Snakemake 9.0.

Jens and Johanna visited the workshop Data Structures in Bioinformatics in Pisa and presented their work on probabilistic filters.
Jens gave a talk about Blocked Bloom filters with Choices. This approach improved the FPR of Blocked Bloom filters by a more balanced load per block.
Johanna gave a talk about Better Cuckoo filters, introducing a windowed memory layout to increase the fill rate of Cuckoo filters.

We had wonderful days at GCB 2024 in Bielefeld. The group was very active:

Jens presented his paper “Swiftly Identifying Strongly Unique k-Mers” at the Wonderful Algorithms in Bioinformatics conference.

We are happy to announce that Sven, Jens and Johanna are organizing a whole day tutorial at the ISMB 2024 in Montréal. We will give an introduction on Just-in-time compiled Python for bioinformatics research.

We are happy to announce that Sven, Jens and Johanna are organizing a half day workshop at the GCB 2024 in Bielefeld. We will give an introduction on Just-in-time compiled Python for bioinformatics research.

At the graduation ceremony of the SIC, Johanna was awarded the Günther Hotz Medal for an outstanding Master’s degree in Bioinformatics.

Sven, Jens and Johanna visited the workshop Data Structures in Bioinformatics in Montpellier. Johanna gave a talk about EpiSegMix, a new tool for chromatin segmentation based on epigentic marks using HMMs.
Andrea Volkamer, Sven Rahmann, and Alexey Gurevich hosted an inauguration / Christmas party at the Center of Bioinformatics.
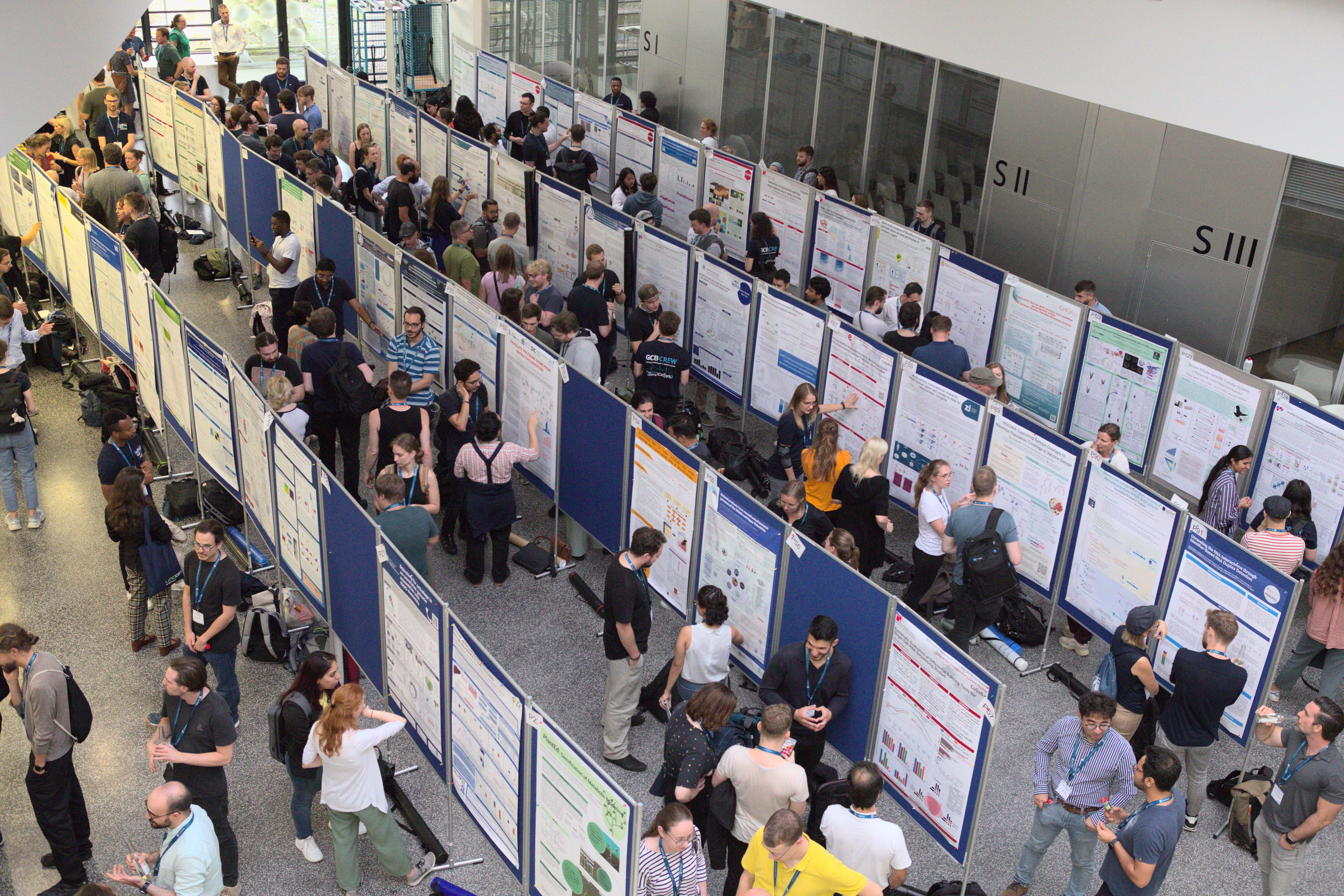
The group visited the German Conference on Bioinformatics (GCB) in Hamburg. Sven co-organized the workshop Education in Bioinformatics. Johanna, Vu Lam and Andre presented posters about their current work. Jens gave a talk about the new Xengsort version.

Vu Lam Dang and Jens Zentgraf participated in the CPM summer school at the École normale supérieure in Paris. Vu Lam presented a poster about his work on factorization of binary matrices and the possibility of using them to calculate polygenic risk score. Jens presented a poster about the possibility to combine super-k-mers and multi-way bucketed parallel Cuckoo hashing.

From 14.06. to 16.06 Andre Holzer, Sven Rahmann, Johanna Schmitz, Johannes Schreieck and Jens Zentgraf participated in the HLA-Workshop by the Stefan Morsch Foundation. Andre gave a talk about the Opportunities and Challenges of Long-Read Sequencing.

Sven Rahmann and Jens Zentgraf visited the group of Jan Fostier in Ghent. We had an interesting exchange about search schemes, k-mers and different applications.

We visited the Stefan Morsch Foundation and had the opportunity to visit the laboratories and look at the procedure. We discussed the possibilities, difficulties and advantages of long read sequencing in HLA typing.

On 01.04 we could welcome Johanna Schmitz as a new member of the group. She has already written her master thesis entitled “Multivariate Hidden Markov Models with Flexible Distributions for Chromatin–State Discovery” in the group and is working on:
Algorithmic Bioinformatics, SIC, Saarland University | Privacy notice | Legal notice